Neuroevolutionary Wordle - Garbage Collection and the Older Generation
The Problem - Increasing the Population Size
When breeding the next generation in a genetic algorithm, the elder generation provides its genotypes for recombination and mutation. This means there is a time when two generations of genotypes are needed in memory.
To squeeze the most power out of a domestic GPU, we may want to find a more efficient way of managing that. Is there a way of producing the second generation while the first still exists, without using 2x the memory of a single generation?
Other Measures Used
Other measures implemented or considered have included:
- Decreasing weight precision in the underlying neural nets being bred from fp32 to fp16;
- Dropping the number of 5-letter words the models actually know from 15k to 5k;
- Inventing a novel solution to late-stage action injection in output embeddings - which is Wordle-specific in its nature - to allow for smaller models and larger populations before they’ve learnt all the 5-letter words;
- Potentially attempting to cause a smaller population to behave like a larger one, through spatially-aware training data sharding/sampling, to improve genetic diversity.
But, as is always the case when it comes to evolution, it’s never enough…
Introducing a Shared Genotype Buffer for Genetic Algorithms
The first thing to say is that this did not start out as a deliberate attempt at a garbage collector. The language has been changed after helpful feedback from a reviewer. I have come to accept that it is a garbage collector.
Specifically, the code now has a reference counting, compacting garbage collector.
That being cleared up, some changes to the memory structure are needed. Specifically, imagine we replace our two concurrent generation buffers (in VRAM within our CUDA code) with a single slab. The slab can contain any organisms, of either generation. Their genotypes need only be placed in whatever slot is available. A generation is now simply an array of references into the slab, to fixed-width (i.e. fixed ‘stride length’) slots. One genotype gets one slot, and it doesn’t really matter where.
How a Shared Slab for All Genotypes Actually Helps in a GA
You may be wondering how this actually helps. After all, we still have the same number of genotypes to store in memory. We have just made things a little more complicated, without saving any memory yet.
The important thing to understand is that not all of the elder generation are actually needed. The fitness function has computed their evaluations. In some cases, it isn’t looking too good. According to the next generation’s assembly plan, many are destined to die childless, for being bad at Wordle. Darwin never said this system was kind.
In using a garbage collector style ‘reference counter’ approach, we keep a record of how many children a given genotype in the parent generation is still yet to have. This gets decremented when recombination/mutation creates the child genotype. At zero, it indicates that the slot can be freed up. A reference into the slab can be deleted; the old genotype soon to be clobbered by one of its neighbour’s children.
Using reference counter garbage collection in the shared, bi-generational genotype slab, we may make a substantial saving in memory pressure, allowing a larger population to evolve on the GPU.
How Much Memory Does This Save?
Time will tell, but I’m aiming to reduce the size of the shared genotype slab from the twin buffers approach of 2x the size of a population to just 1.4x, and see what happens.
What If OOM?
If the number of slots available isn’t sufficient, a failover to host RAM should take place. This is not to be taken lightly: the warnings must be loud, and any such failover should be treated as a bug. We cannot get through a reasonable number of generations of our genetic algorithm if the CUDA genotype slab runs out of memory, and we have to transfer part of the population over a PCIe bus to process it on-host, then shift it on-device again in time for that generation’s fitness evaluation. Very occasionally failing over to host memory is better than a long GA run crashing, but this cannot be a regular occurrence from one generation to the next. The population size should be decreased until this basically doesn’t happen.
Summary
A single shared genotype slab will allow us to save memory, and therefore go with larger population sizes. The genotype slab will contain members of both the current and next generation. Genotypes will be addressed by offsets or indices into the genotype slab; and a generation is just an array of pointers to the starts of genotypes. When a genotype within the current generation is destined not to be part of a breeding pair, or when it has already been used for recombination and mutation, it is no longer needed. Deleting its pointers and freeing up the slot creates more space, for the storage of a member of the child generation under construction. I’m going to try to get away with using 1.4x the size of a single generation for the genotype slab, failing over to host memory if unavoidable.
Non-Fixed-Width Slots: Making the Memory Allocation More Complicated Still
Why Non-Fixed-Width Genotypes
Just when our storage of genotypes looked as unnecessarily complicated as possible, another issue comes up.
Remember the idea about ‘Injection of New Output Embeddings’? A model has an internal dictionary of 5-letter words, each of which is represented by 26 derived values, and 38 trainable values (which need to go in memory), to make a 64-dimensional vector in its output embedding.
With an internal lexicon of around 5,000 words, these output embeddings weigh in at fully half of the total number of parameters of the model. Making that smaller would allow a larger population size, giving better exploration of the search space of possible Wordle-playing genotypes.
What can be achieved here is a phased curriculum of training data. The idea is simple: run the GA with just 50 possible solutions to a Wordle used in fitness evaluation, and only give the model 50 corresponding output embeddings. Introduce new ‘shards’ of training data, in 50-word batches, periodically during the GA run.
This required a novel, Wordle-specific solution to late output embedding injection. We now have a model whose architecture is:
- An input encoder, used up to 5 times, for each previous guess in a Wordle Puzzle [fixed size]
- A dense trunk neural net [fixed size]
- An output embedding of 64-dimensional vectors representing the words the model knows [grows in size]
Ultimately, the point of this is to allow the population size to be even bigger in early generations, but there is a problem.
This is going to play havoc with our shared, bi-generational genotype slab slot system, if only in those generations where the model size actually grows.
Compacting the Garbage
The 5070 Ti needs a theoretical floor of 36ms for reading and writing 16GB of VRAM. The broader compaction process is probably only needed once per 100 generations. This depends on the training data phased curriculum schedule demands.
In reality, we may spend 100-200ms on compaction, when it is needed. This is acceptable, as it is not every generation.
Here is how it can work:
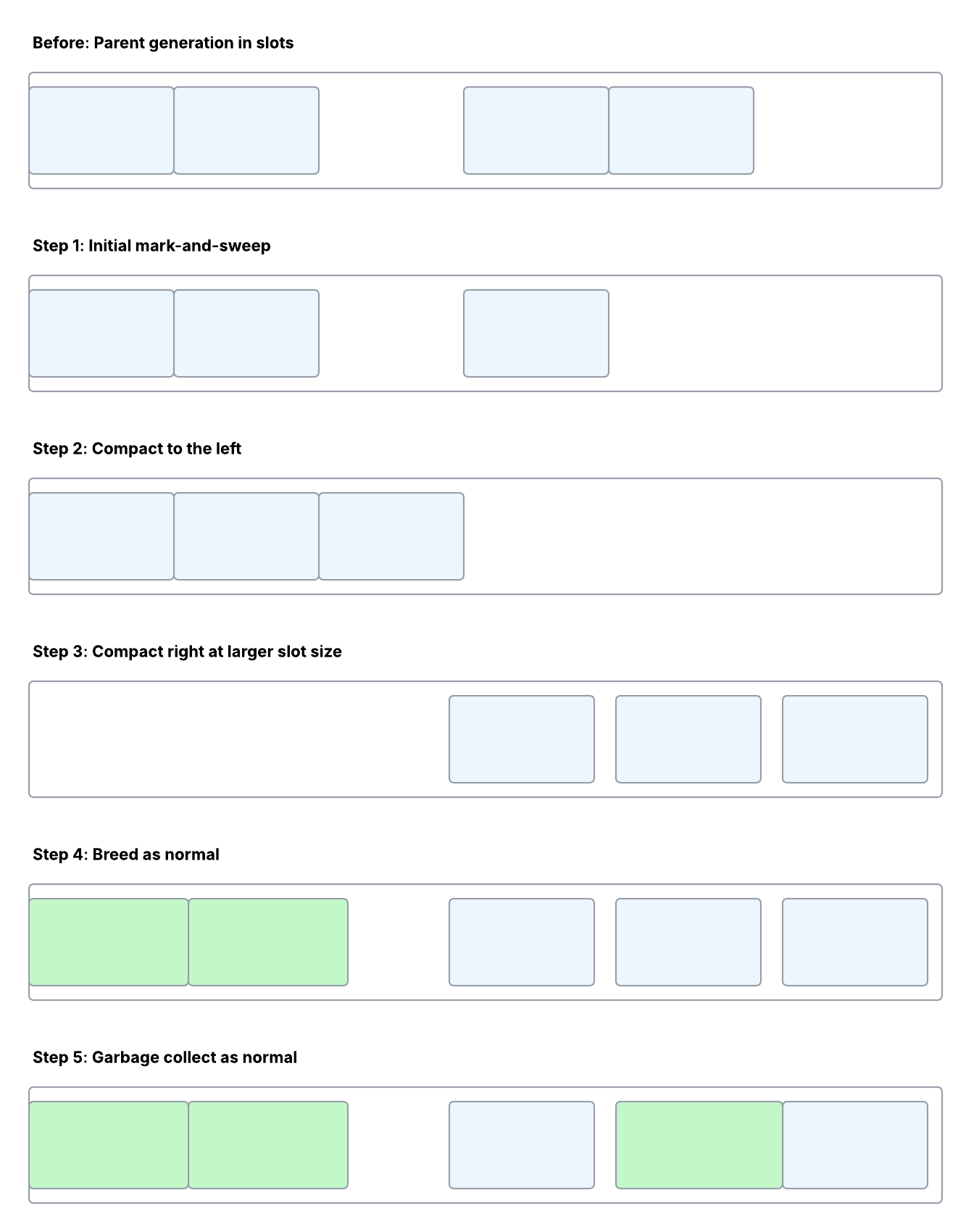
Step 1: Initial Deletions Based on Reference Counter
Any organisms in our population destined not to have offspring may as well get cleared out first, in the normal way.
Step 2: Everything You Own in a Box to the Left
Compact once, using the ‘old’ slot size, to the left-hand side (i.e. lower memory addresses).
Step 3: Expand and Repack Right
Repack, this time using the new slot size. Working right-to-left, move everything to the free space on the right, thereby ensuring that other genotypes don’t get ’trampled’ during repacking. The increased slot size seems likely to be an increment of 3.8kB. The initial slot size for the first generation could be around 300kB, but that remains to be confirmed.
Step 4: Breed as Normal
Recombination and mutation continue as normal.
Step 5: Keep Garbage Collecting
Whenever a parent has discharged all of its duties, according to the next generation’s assembly plan, its slot gets freed up, and a new member of the child generation may take its place
Summary
We can still use a single shared genotype buffer. With a modest amount of reshuffling inspired by Javascript and Ruby style garbage collectors, we can accommodate a step-change in the necessary slot size, while maintaining the advantages of a shared genotype buffer system.
When new 5-letter words are introduced to the models, their genotype needs to grow. The population size can shrink, to maintain constant memory requirements. This way, we are able to enjoy a larger population size in earlier generations, squeezing a little extra exploration of the fitness landscape out of a genetic algorithm running on a domestic GPU.